 Resource Library
/
Accurate Mass: The best solution for metabolite identification in discovery, development and clinical applications.
Resource Library
/
Accurate Mass: The best solution for metabolite identification in discovery, development and clinical applications.
white-paper
Accurate Mass: The best solution for metabolite identification in discovery, development and clinical applications.
In this paper, we review appropriate definitions and describe the theoretical basis of why accurate mass approaches offer a significant advantage over nominal mass approaches in the arena of qualitative analysis and, specifically, metabolite identification.
Register to gain access to gated resources.
"*" indicates required fields
Resources to Consider


webinar
Rapid, accurate and highly customizable assay platforms for...
The diagnosis and treatment of human diseases and the development of new drugs increasingl...

journal
Accurate determination of drug-to-antibody ratio of interchain...
Accurate determination of the drug-to-antibody ratio (DAR) of interchain cysteine-linked a...

webinar
BSEPcyte® and MDR3cyte®: Innovative Solutions for Investigating...
Drug-induced liver injury (DILI) accounts for >50% acute liver failures, and is the leadin...

webinar
Strategies and Solutions for Early Phase Formulation Development and...
In this webinar, we discuss the strategies to overcome challenges during formulation devel...

poster
Challenges and Bioanalytical Solutions of Ultra-Low Sensitive 4-in-1...
Achieving ultra-low pg/mL sensitivity, this innovative 4-in-1 hormone biomarker assay demo...

infographic
CMC-Product Development & Manufacturing Integrated Solutions
Frontage provides integrated solutions to our clients including Formulation, Product Devel...
ebook
Bioanalysis of antibody-drug conjugates (ADCs) by LC/MS: challenges...
In this eBook, we explore the various ways LC-MS is utilized for the bioanalysis of ADCs a...

journal
Drug-drug Interaction Study of CYP3A4 Inhibitors with Colchicine Oral...
The primary objective of the study was to compare the pharmacokinetic (PK) variables of pl...
journal
Relative Bioavailability Study of Colchicine Oral Solution and...
In this study, we evaluate the pharmacokinetics of colchicine oral solution, and determine...

journal
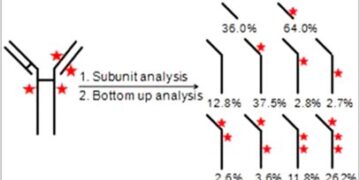
Characterization of Positional Isomers of Interchain Cysteine Linked...
Interchain cysteine linked antibody–drug conjugates (ADCs) are emerging therapeutic prod...

poster
Comparison of the Quantitative Measurement of Albumin in Human Plasma...
Human serum albumin (HSA) is the most abundant protein in human blood plasma. This poster ...

journal
An Ultrasensitive LC-APPI-MS/MS Method for Simultaneous Determination...
An Ultrasensitive LC-APPI-MS/MS Method for Simultaneous Determination of Ciclesonide and A...

