Genomics
Metagenomics & Microbiome
Solving development problems of probiotics and prebiotics
Frontage offers a comprehensive portfolio of genomics services to meet the needs of our clients.
We provide solutions to our clients from Pre-Clinical & Discovery through to Clinical Trial & CDx Support. Frontage uses advanced technologies and state-of-the-art equipment to provide fast, accurate, and customized results you can trust.
Comprehensive Genomics Services
Frontage offers end-to-end genomics services, from nucleic acid isolation through data analysis and interpretation, to provide manageable datasets and comprehensive results packages.
See here for a full list of our solutions and technologies.
- Genomics Services for Oncology
- Cell & Gene Therapy Support
- Genotyping / Variant Detection
- Metagenomics / Microbiome
- Pre-Clinical & Discovery
- Clinical Trial & CDx Support
- Genomics Solutions & Technologies
- Bioinformatics & Statistical Analysis
Rapid Turn-Around Times
At Frontage, we pride ourselves on the ability to offer rapid turn-around times for all genomics services. Our workflows are optimized for high-throughput samples processing to accommodate a large number of samples in a short amount of time.
Meeting Regulatory Standards
Our genomics services laboratory is CLIA-certified and CAP-accredited and meets the GLP regulatory standards.
Frontage offers pre-validated NGS-based LTDs for oncology and pharmacogenomics that are CAP-accredited. Custom NGS, qPCR, or droplet digital PCR assays can be set up and validated under GLP or CLIA.
Frontage offers advanced next-generation sequencing technologies for microbiome profiling for patient stratification, clinical trial support, and basic research. Our workflows are optimized for processing various sample types, allowing for the study of microbial communities from the human gut, lungs, skin, and various other environmental niches.
The end-to-end metagenomics services offered at Frontage conclude with comprehensive metagenomic analyses for identifying taxa present in the sample, elucidating taxonomic diversity and make-up, as well as phylogenetic relationships of organisms of the microbial community.
Interactive Metagenomics Report

Microbial profiling using 16s rDNA sequencing
Deep sequencing of 16s rDNA amplicons allows for the discovery, identification, and quantification of diverse prokaryotic organisms living in a community. Frontage offers sequencing of the 16s rDNA with primer sets that span both V3 and V4 hypervariable regions which have been recommended by Illumina (BK primer set), or an alternative set that spans V4 only and which has been utilized by the Earth Microbiome Project (PA primer set), the latter having been optimized for detection of Archaea. Both primers sets have been validated in Frontage’s laboratory. Amplicons are then sequenced with the latest Illumina instruments and technology.
Deep sequencing of 16s rDNA amplicons with BK or PA primer sets
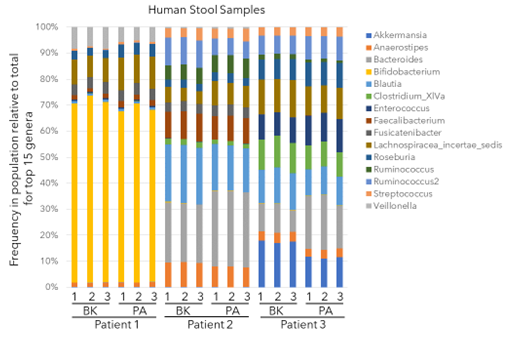
The stacked bar chart above shows the frequency of the top 15 genera relative to the total for 18 DNA sequencing libraries from 3 different patient stool samples. Genomic DNA was isolated from each of the 3 patient stool samples in triplicate. The 16s rDNA from each set of triplicates was amplified with both the BK primer set and PA primer set, for a total of 18 DNA sequencing libraries. The results show consistency in identifying and quantifying genera between replicates and across both primer sets. The largest variation in the gut microbiota is observed between different patients.
For more information on this, read our white paper: 16S rDNA Amplicon Sequencing of Mock Microbial Community Standards.
Custom metagenomics analysis to meet your needs
In addition to our standard 16s rDNA profiling, Frontage can also implement custom primer sets or whole genome sequencing for clients with unique experimental goals.
Metatranscriptomics
Frontage’s metatranscriptomics services facilitate gene expression studies on a global scale, revealing the transcriptionally active portion of the microbiome, i.e., the portion that contributes to the functional capacity of the microbiome. Our metatranscriptome services are offered for all stages of your microbiomics project. We perform RNA extraction, library preparation, sequencing, and analysis of your complex metatranscriptomic data including interactive taxonomic classification reports, as well as gene-level expression analysis and visualization.
Metatranscriptomics can be used in multiple areas of research:
- Determine drug treatment responses with measurable changes in microflora composition and gene expression
- Microflora RNA signatures can be associated with specific-disease subtypes and enable patient stratification, including identification of non-responders
- Study induction or repression of specific metabolic pathways in response to changes in diet or during disease development
- General mechanism of action studies
- Improve diagnostics for infectious disease status and susceptibility
- Study interactions between symbiotic bacteria and host
DNA Isolation optimized for different organisms
Frontage’s metagenomic analysis service starts with DNA isolation using methods designed for uniform and efficient lysis of many different organisms (e.g. Gram-negative/positive bacteria, fungus, protozoans, and algae) from a variety of sample inputs.
Accepted Sample Types
- Stool/feces
- Tissue biopsies
- Cells
- Saliva
- Bronchial lavage
- Sputum
- Buccal swabs
- Soil samples
- Water samples
Resources To Consider

Metagenomics Services
Microbiome Services Fact Sheet
Genomics: A Customized NGS Sequencing


