case-study
High Throughput Sample Analysis Using Gyrolab
In this case study, Frontage successfully completes the method development and validation under the Sponsor's pressing timelines, while providing high quality, and reproducible data with an ISR passing rate of over 90%.
Register to gain access to gated resources.
"*" indicates required fields
Resources to Consider


poster
WRIB 2022: Optimization and standardization of RNA-seq operation to...
We standardized an RNA-seq operation method under Good Lab Practice (GLP) standards, which...

poster
Optimization and standardization of RNA-seq operation to get...
To ensure good data yield, there are challenges encountered during NGS operations, especia...

poster
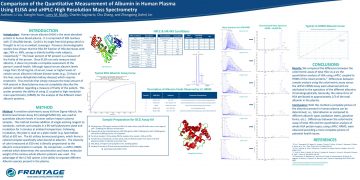
Comparison of the Quantitative Measurement of Albumin in Human Plasma...
Human serum albumin (HSA) is the most abundant protein in human blood plasma. This poster ...
poster
Utilization of High Sensitivity Proteomics NULISA HT Panels for...
This first-ever reported study demonstrates the rigorous qualification of NULISAseq high-t...

journal
Gemogenovatucel-T Advantage in Clonal Tumor Mutation Burden–High...
In a recent clinical study, Gradalis’ Vigil showed a significant survival benefit in cTM...

poster
WRIB 2024: High-quality RNA-seq Library Preparation from...
Here, we standardized the RNA extraction method from FFPE slides and an RNA-seq operation ...

poster
IN Vitro Assessment of the inhibition of Human Organic Anion...
The present study was aimed to validate this alternate approach to assess inhibition of or...

poster
Validation of Ultra-high Sensitive SIMOA Assays for the Quantitation...
In this poster, we validate ultra-high sensitive SIMOA assays for the successful quantitat...

webinar
Rapid, accurate and highly customizable assay platforms for...
The diagnosis and treatment of human diseases and the development of new drugs increasingl...

webinar
Biomarker and Alzheimer’s disease: role of high sensitivity...
It has been said that the diagnosis of Alzheimer's disease enters the era of biomarkers. T...

journal
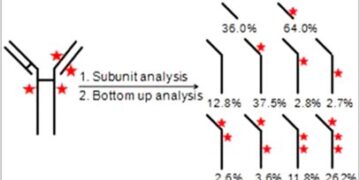
Characterization of Positional Isomers of Interchain Cysteine Linked...
Interchain cysteine linked antibody–drug conjugates (ADCs) are emerging therapeutic prod...